Single-molecule technique to explain thermodynamic behavior of complex biomolecular systems
Most thermodynamic measurements of binding reactions rely on the validity of the law of mass action and the assumption of a dilute solution. Yet, important biological systems such as allosteric ligand-receptor binding, macromolecular crowding, or misfolded molecules may not fulfill these assumptions and may require a particular reaction model. Today in an article published in Science journal by a team of the University of Barcelona, researchers have determined thermodynamic properties in biomolecular complex systems such as binding energies, chemical selectivity, and allostery of nucleic acids and peptides in a model-independent way.
The study has been fully carried out at the Small Biosystems Lab of the University of Barcelona, led by Fèlix Ritort, professor at the Department of Condensed Matter Physics of the Faculty of Physics of the UB, and has counted with the participation of the researchers Joan Camuñas and Anna Alemany.
“This research extends the use of fluctuation theorems beyond unimolecular folding reactions, bridging the thermodynamics of small systems and the basic laws of chemical equilibrium. Furthermore, a similar approach could be used for more complex RNA-protein and protein-protein interactions,” explains Felix Ritort, .
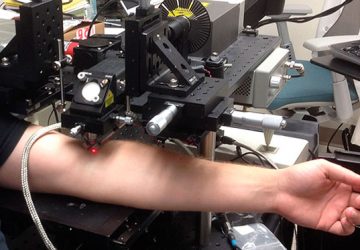
Binding energies are key quantities that determine the fate of intermolecular reactions and force techniques such asoptical tweezers applied to single-molecules, which can be used to pull on individual ligand-DNA complexes, allowing the detection of binding events one at a time.
“We introduce a fluctuation theorem for ligand binding (FTLB) that allows us to directly extract binding energies of bimolecular or higher order reactions from irreversible work measurements in pulling experiments” explains Ritort.
Single-molecule manipulation is particularly suited to observe the formation of misfolded structures, but there is a lack of methods to characterize these systems. By applying the FTLB, it was possible to extract the binding energy from these kinetically stabilized non-native structures.
“By using a DNA hairpin with two binding areas/spots separated by 4 pairs of bases, we observed that the formation of a misfolded structure consists of two short hairpins series” says Ritort. Using different biomolecular systems of increasing complexity, we provided with a single-molecule verification of the law of mass action and showed how the FTLB can account for mass exchange between a molecular system and the environment. “The FTLB provided with a direct experimental measurement of binding energies without assuming any model or reaction scheme, which is particularly useful in cases where the law of mass action doesn’t work ,” concluded Ritort.
source : www.sciencedaily.com