Molecular pathways revealed could identify potential drug targets for restoring hearing
Multiple new genes involved in hearing loss have been revealed in a large study of mouse mutants. The new genes reveal the metabolic pathways and regulatory processes involved in hearing.
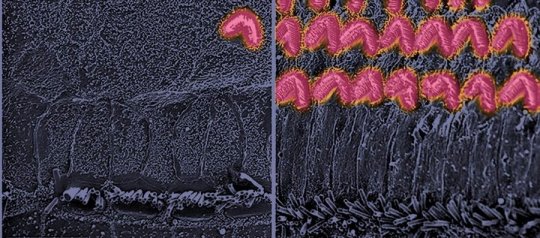
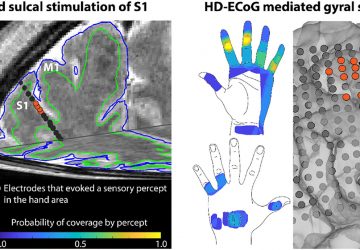
Credit: Ingham et al. (2019) PLOS Biology DOI: 10.1371/journal.pbio.3000194
Multiple new genes involved in hearing loss have been revealed in a large study of mouse mutants by researchers from the Wellcome Sanger Institute and King’s College London, and colleagues. The new genes identified reveal the metabolic pathways and regulatory processes involved in hearing.
The study, published today (April 11) in the open-access journal PLOS Biology, helps to understand the underlying biology of deafness, and also provides a rich source of therapeutic targets for the restoration of hearing.
Progressive hearing loss with age is extremely common in the population, leading to difficulties in understanding speech, increased social isolation and associated depression. It can often be inherited, but so far very little is known about the molecular pathways leading to hearing loss, hampering the development of treatments.
To identify new molecules involved in hearing loss, the researchers took a genetic approach and created 1,211 new mouse mutants. They screened each of these mice using a sensitive electrophysiological test, the auditory brainstem response, to find out how good their hearing was.
This large-scale screen of targeted mouse mutants identified 38 genes involved in hearing loss in the mice, that had not been previously suspected to be involved in hearing.
The researchers also analysed human DNA data to ask if any of these 38 genes discovered in mice were associated with human adult-onset hearing loss. They found eleven of these 38 genes were significantly associated with hearing ability in the UK population. Furthermore one gene, SPNS2, was associated with childhood deafness.
Some of these genes revealed molecular pathways that may be useful targets for drug development.
Dr Chris Lelliott, an author from the Wellcome Sanger Institute, said: “This is the first time that a study of this scale has looked at levels of hearing and different types of hearing loss in mouse mutants and shows the power of large genetic screens. Only a handful of genes have previously been linked specifically to age-related hearing loss in adults, now our study adds many more potential new genes to follow up.”
Further analysis of the genes identified, and the many different mechanisms within the ear that were revealed by the mutations, suggested that hearing loss is an extremely varied disorder and may involve as many as 1,000 genes.
Dr Selina Pearson, from the Wellcome Sanger Institute said: “This study is giving a huge insight into the complicated biology of hearing loss, and shows that because of all the different genes and pathways found, there won’t be a single ‘magic bullet’ to stop all age-related deafness. This emphasises the value of mouse studies for identifying genes and mechanisms underlying complex processes such as hearing.”
The study findings suggest that therapies may need to be directed at common molecular pathways involved in deafness rather than individual genes or mutations.
Prof Karen Steel, senior author on the paper from the Wellcome Sanger Institute and King’s College London, said: “Several of these new mouse mutant lines showed normal development of hearing followed by later deterioration, suggesting the genes involved are good candidates for human age-related hearing loss. Our next step is to find out if we can influence the molecular pathways involved to slow down or stop the progression of hearing loss.”
source: www.sciencedaily.com