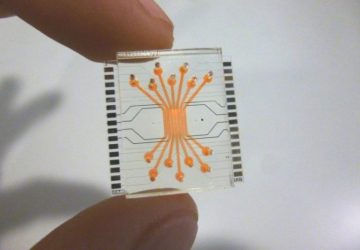
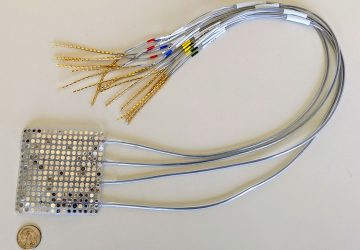
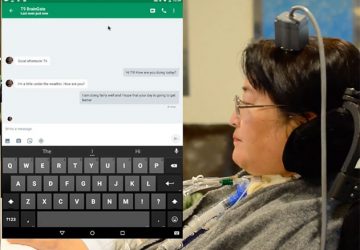
Fluorescent tagging system can expedite the process of designing genes and personalizing medicine.
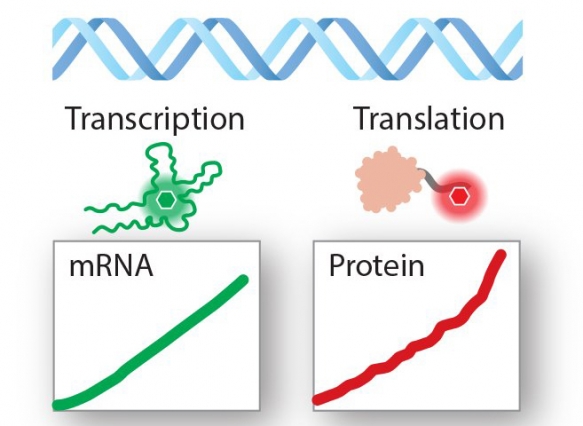
Image courtesy of MIT Lincoln Laboratory
Builders of genetic circuits face the same quandary as builders of digital circuits: testing their designs. Yet unlike bioengineers, engineers have a simple and universal testing tool — the multimeter — that they can touch to their circuit to measure its performance. “There’s nothing remotely like this in bio,” says Peter Carr, a synthetic biologist in MIT Lincoln Laboratory’s Bioengineering Systems and Technologies Group.
That was, until recently. Carr and researchers in his group have developed a system that they liken to a “biomultimeter.” The system, called PERSIA, uses fluorescent labeling to illuminate different parts of a genetic circuit and allows researchers to measure biological functions — including transcription, translation, and other enzyme activities — in vitro in real-time. A paper describing this work was published online in the journal ACS Synthetic Biology.
“We now have a way to quickly test new genetic code designs in specific genes. We can accelerate how we ask and answer questions with DNA,” says coauthor David Walsh.
Fluorescent labels
For synthetic biologists to engineer cells that work how they want them to — say, to be immune to a virus — they must be able to tell the cell which genes to express. To do this, they use synthetic DNA to program genetic circuits that control the cell’s behavior. By measuring transcription (the process by which a gene’s DNA is copied into RNA) and translation (the process by which that RNA is read to produce proteins) bioengineers can know how well a circuit is working.
Proteins such as green fluorescent protein (GFP) are already a favorite testing tool for gene expression. With such methods, a gene that carries instructions to produce fluorescent pigments can be fused to a gene in a DNA sequence that will produce a protein of interest. The GFP glow lets biologists know that this protein is being produced.
While GFP has been game-changing for bioresearch, it has its limitations. For one, fluorescent proteins are large, comprising over 200 amino acids that eat up the finite resources in an in-vitro test and may interfere with the natural function of the protein of interest. They also take time to emit their signal; thus, GFP is a reporter of the past, and when things don’t go as planned, it’s unclear what went wrong and when.
PERSIA, on the other hand, uses small fluorescent tags appended to the researcher’s protein of interest. Because PERSIA’s tags are so small, they can be attached to proteins anywhere within a genetic circuit, allowing for the study of different parts of it. They also emit faster than GFP by several minutes, more representative of live rates of transcription and translation.
“We are replacing GFPs with something smaller, that is easier to move around and reconfigure, and measuring fluorescent signals as these processes occur,” Carr says.
PERSIA stands for PURExpress-ReAsH-Spinach In-Vitro Analysis and uses the off-the-shelf products listed in its name. PURExpress is the cell-free “soup” that houses the experiments; this type of in-vitro environment allows for the study of specific biological processes apart from a full living organism.
Spinach is the name of the RNA tag that is fused to a gene. It will fluoresce green when the gene’s DNA is transcribed into RNA and as it reacts to a chemical reagent in the PERSIA mixture. A second, peptide tag is also attached to the gene. This tag will fluoresce red as it is translated at the end of a protein and upon reacting with another chemical reagent in the PERSIA mixture called ReAsH.
During an experiment, all of these ingredients — the tag-encoding DNA sequences and the reagents — are mixed together. Fluorescence is monitored. The amount and the activity of RNA and proteins, the products of transcription and translation, are essentially immediately known.
Genetic and clinical designs
Because of its swift fluorescence, PERSIA can accelerate the evaluation of genetic code designs and act as a screening tool for obvious failures. “You can project what is going to happen,” says Scott Wick, first author of the paper. “Weeks if not months of time are saved downstream for prototyping, testing different designs out and doing many more tests before you get into exploring these engineered codes in vivo.”
The developers tested PERSIA by using it to screen their own redesigns of the genetic code of E. coli. Changes in the ReAsH fluorescent signal showed that one of their redesigns caused translation rates to plummet. They then studied which genetic changes specifically might be responsible for this defect and constructed synthetic DNA to evaluate their best guesses. With only a few DNA sequence modifications and tests with PERSIA, their revised gene design was as productive as the original E. coli gene.
“PERSIA let us quickly troubleshoot how protein production had gone way off and revealed how we could bring it back to normal levels,” Carr says.
Beyond its use for reporting on transcription and translation, PERSIA is also valuable for studying how a protein is working. For example, the team extended PERSIA to monitor the activity of the enzyme HIV-1 protease, which matures proteins in the HIV virus and is one of the main targets for anti-HIV drugs. They reconstructed clinical variants of the HIV-1 protease gene known to demonstrate resistance to these drugs. They then added a product to PERSIA that fluoresces with HIV-1 protease activity, using the results to monitor the effects of different anti-HIV drugs on each variant. The variants’ resistance patterns as determined from the PERSIA tests were consistent with their known resistance patterns documented in the Stanford HIV Drug Resistance Database.
“When you think about all of the genetic diversity that a viral infection has, this constellation of variants, you want a quick way of testing that diversity for choosing the best drug therapies,” Carr says. “We can do these resistance tests with PERSIA, many of them, and quicker than was previously possible, and hope to use it for giving patients personalized drug regiments. This way the patient isn’t the experiment.”
Similarly, the PERSIA tool could be applied to health security, allowing researchers to respond to new and potentially devastating diseases by rapidly screening the effectiveness of available medicines.
Microfluidic scale
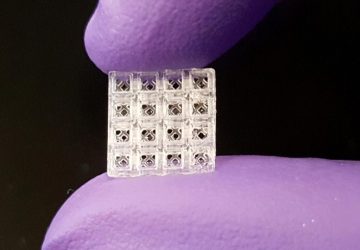
The research team has been implementing PERSIA at the scale of microliters, in standard multiwell plates. But they also demonstrated the potential for miniaturizing the approach in a microfluidic device, at the scale of nanoliters. They designed a microfluidic device with 96 reactors and rows of valves that automatically control the flow of fluids through the device. As the reagents used in PERSIA can be costly, microfluidics provide a way of performing hundreds of tests simultaneously while using up only trace amounts of these chemicals. Building microfluidic devices to include even larger numbers of parallel reactors is one of the team’s next goals.
Another hope is that other scientists will find PERSIA as useful as it has been for the laboratory’s work. The system came together over five years, during which the team noticed that they could use this same, simple method for answering many different questions about their DNA-based designs.
“PERSIA sets a foundation for quickening the process to scientific discovery,” Wick says. “That’s what I’m most excited about.”
source: www.news.mit.edu